Welcome to 2SigFinder!
Genomic island (GI) is a cluster of genes in prokaryotic genomes that have probable horizontal origins. These genetic elements have been associated with rapid adaptations in prokaryotes that are of medical, economical or environmental importance, such as pathogen virulence, antibiotic resistance, symbiotic interactions, and notable secondary metabolic capabilities. The recent development of their detection methods has led to significant advances in our understanding of microbial evolution and function. Despite these advances, several challenges still exist regarding the use of multiscale features, detection algorithm and boundary identification.
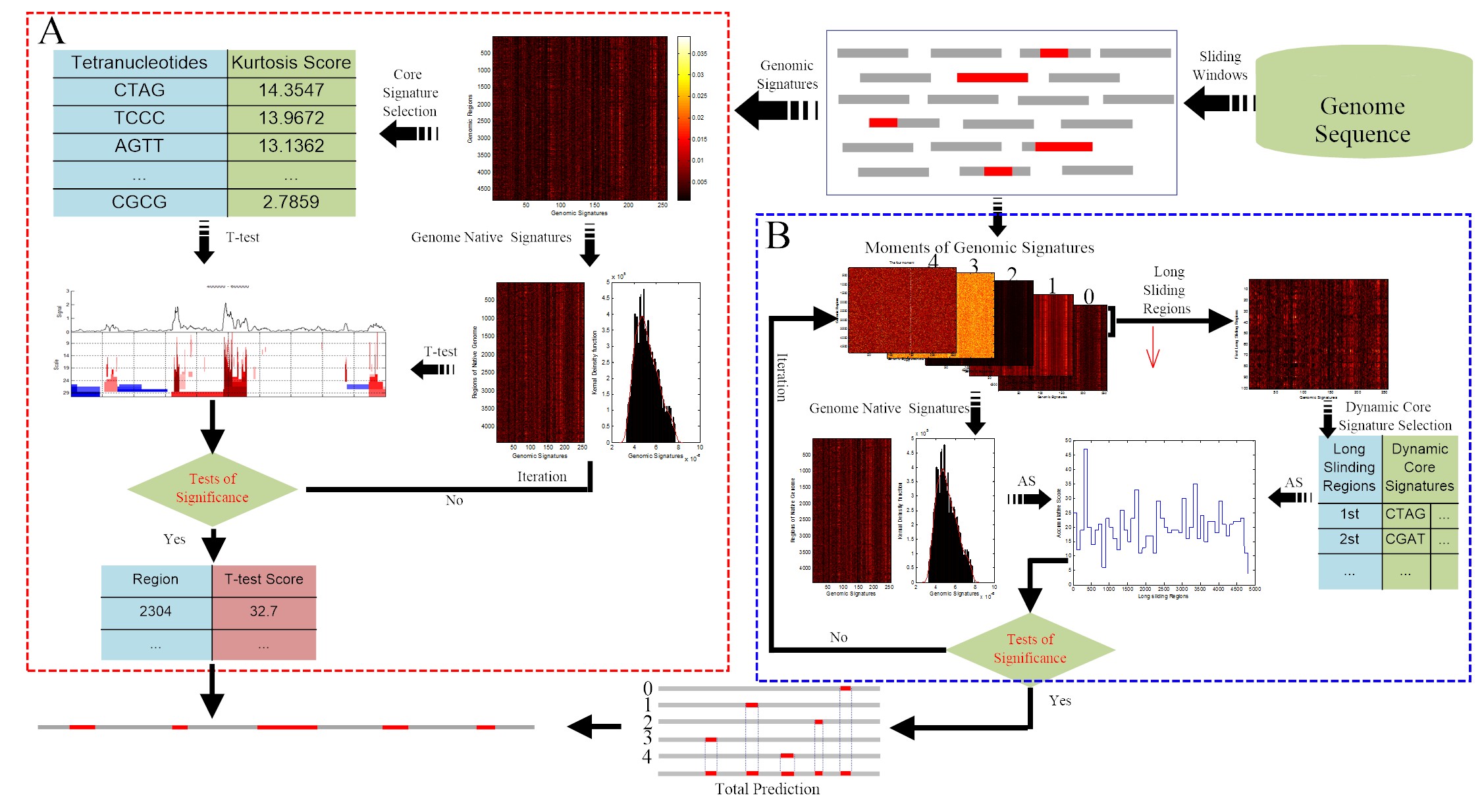
2SigFinder is a novel prediction method, the first combined use of small-scale and large-scale statistical testing for genomic island detection. 2SigFinder was tested by genomic island boundary detection and identification of genomic islands or functional features of real biological data. We also compared the proposed method with the comparative genomics and composition-based approaches. In simulations with alien frag-ments from real genomes, the CG-MJSD searches the maximum of the MJSD score in terms of the GC-content bias, making it much faster in the boundary detection of the genomic islands while main-taining a similar error rate. From real biological data, 2SigFinder identified genomic islands from a sin-gle genome and reported robust results across different experiments, without annotated information of genomes or prior knowledge from other datasets.